Welcome to MetaDraft
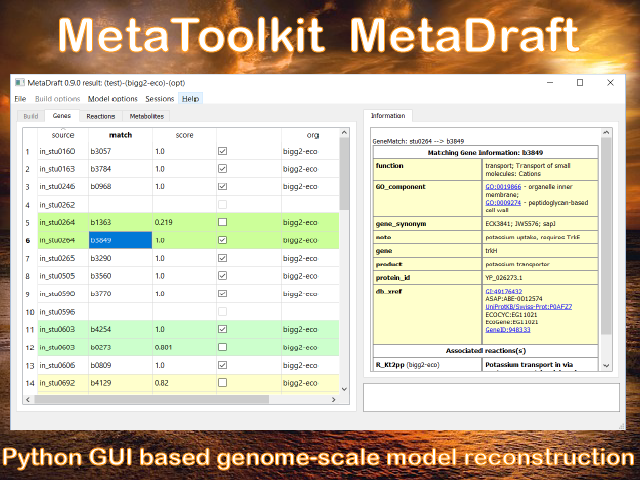
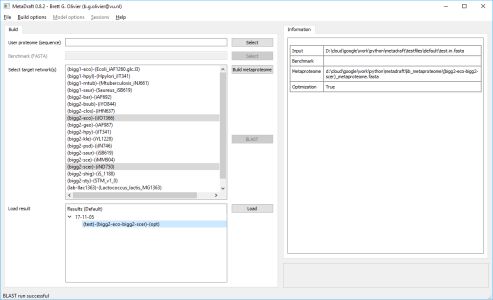
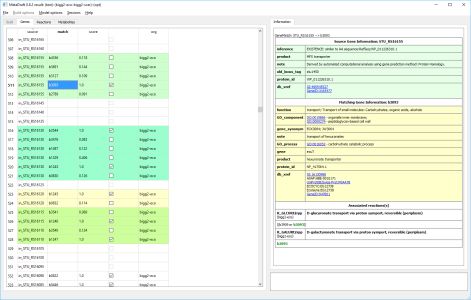
MetaDraft 0.9.2 has now been released. MetaDraft is a full featured GUI based platform for the reconstruction of genome-scale metabolic models. Utilizing a continuously updated, user extendable database of template models MetaDraft provides a stable yet flexible reconstruction workbench for a wide variety of users.
MetaDraft supports the latest Systems Biology standards including SBML Level 3 FBC, Groups, annotations and COMBINE OMEX archives.
MetaDraft forms part of CBMPy MetaToolkit developed in the Systems Bioinformatics Group at the Vrije Universiteit Amsterdam.